Publications
* = equal contribution
' = co-corresponding
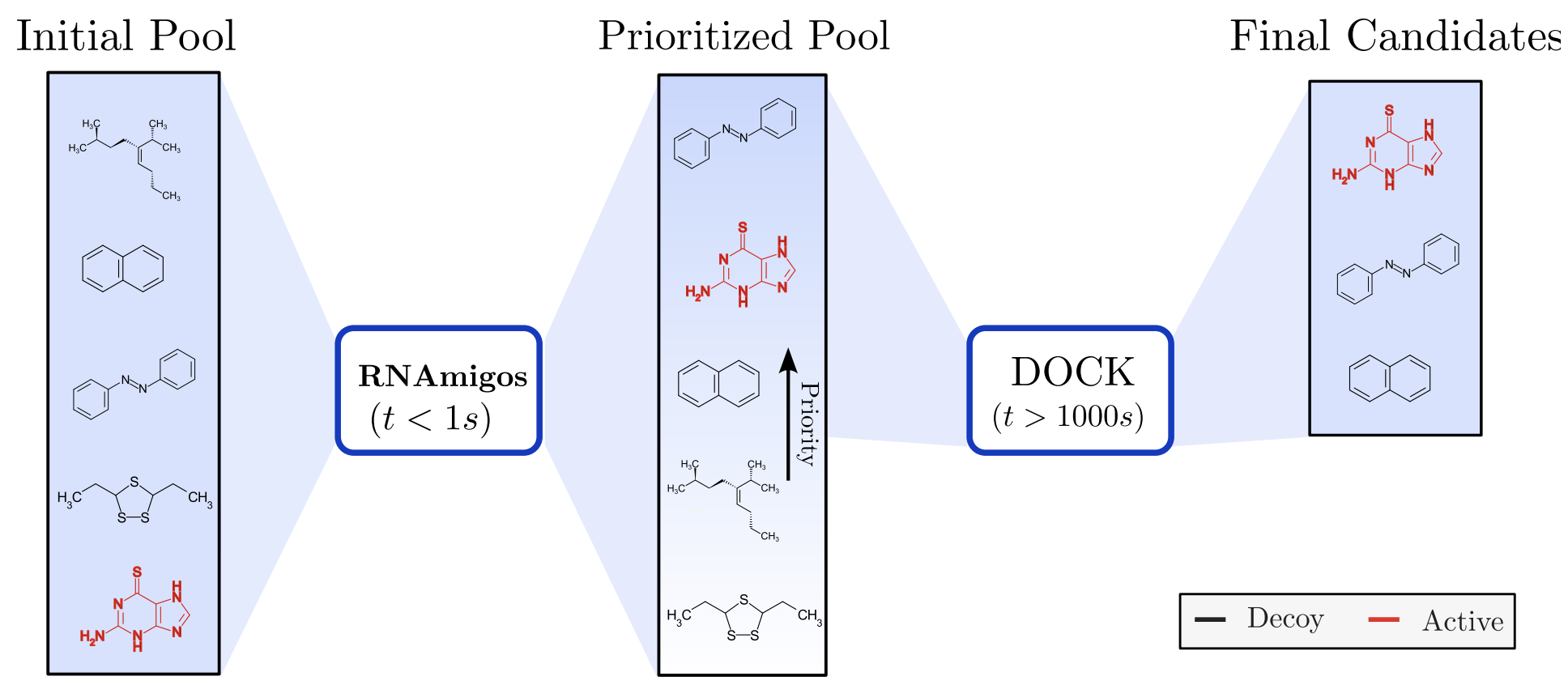 |
RNAmigos2: accelerated structure-based RNA virtual screening with deep graph learning Juan G. Carvajal Patiño*, Vincent Mallet*, David Becerra, L. Fernando Niño V, Carlos Oliver', Jerome Waldispuhl' Nature Communications 2025 (preprint) (article) |
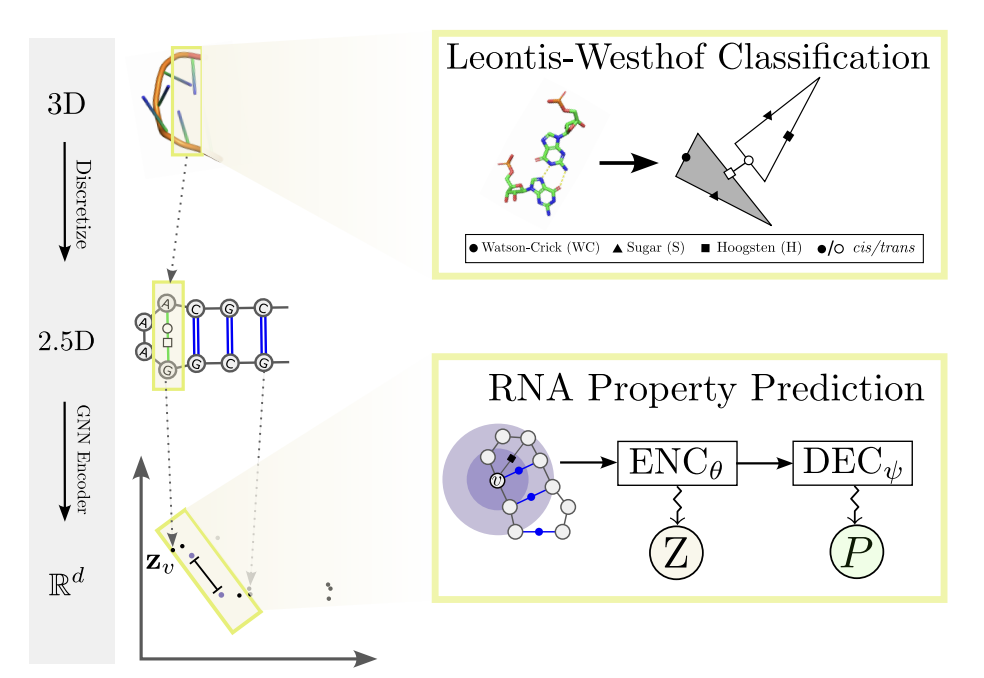 |
3D-based RNA function prediction with rnaglib Carlos Oliver, Vincent Mallet, and Jerome Waldispuhl arXiV (2024), to appear as book chapter in Springer Nature: Methods in Molecular Biology (preprint) |
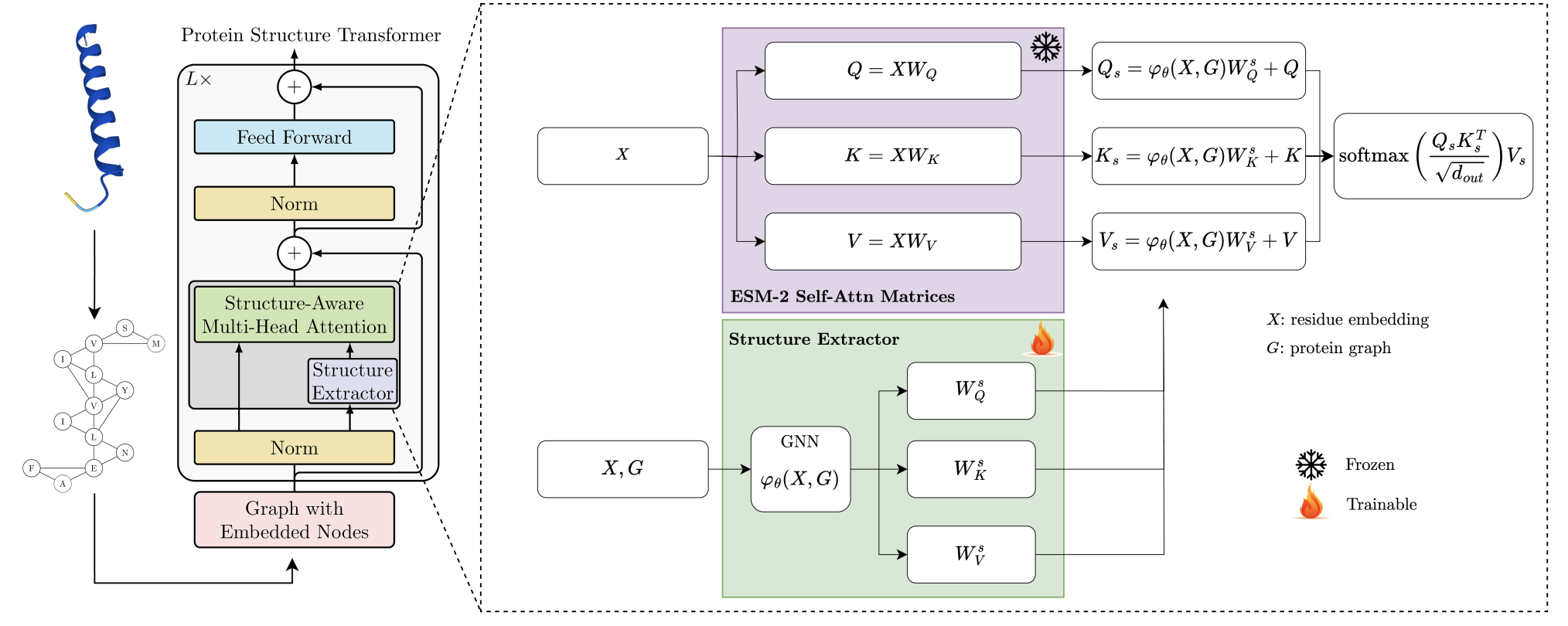 |
Endowing protein language models with structural knowledge Dexiong Chen*, Philip Hartout*, Paolo Pellizzoni, Carlos Oliver, and Karsten Borgwardt arXiV (2024) (preprint) |
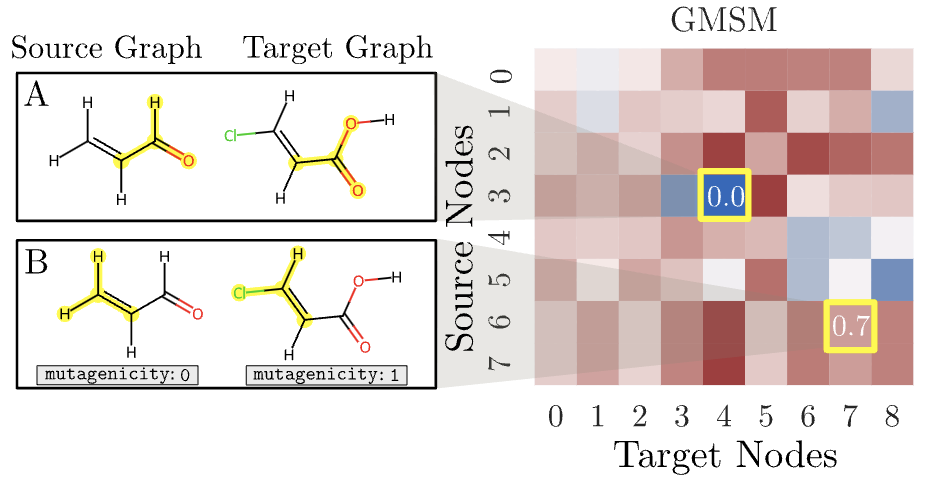 |
Structure-and function-aware substitution matrices via learnable graph matching Paolo Pellizzoni*, Carlos Oliver*, Karsten Borgwardt RECOMB (2024) (proceedings) |
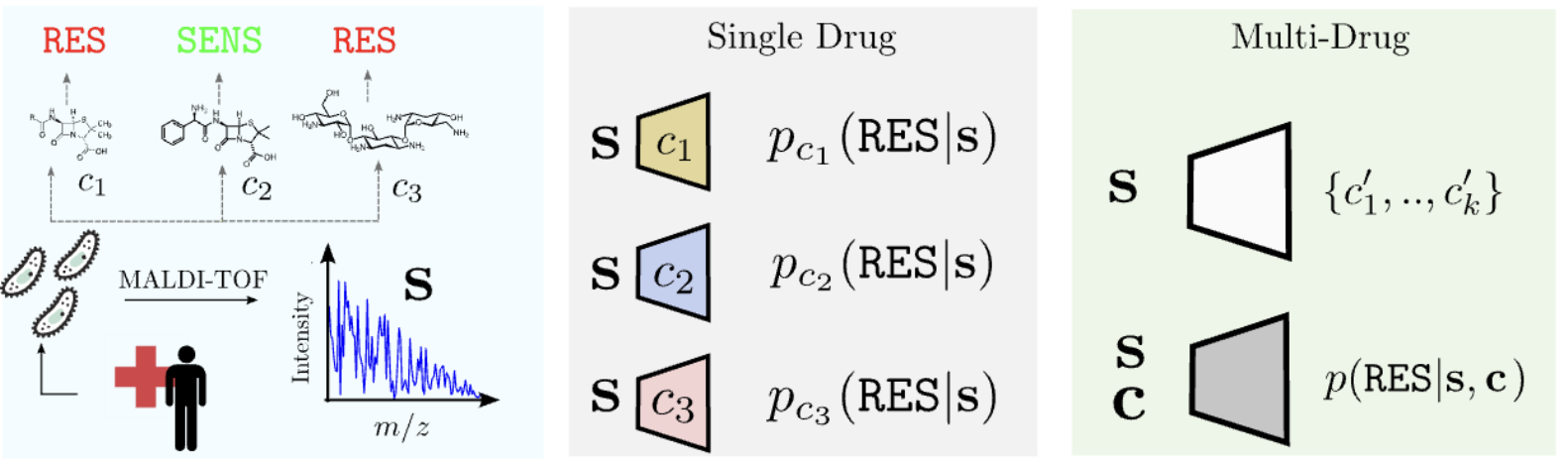 |
Multimodal learning in clinical proteomics: enhancing antimicrobial resistance models with chemical information Giovanni Visonà*, Diane Duroux*, Lucas Miranda, Emese Sükei, Yiran Li, Karsten Borgwardt', and Carlos Oliver' Bioinformatics (2023) (paper) |
 |
ProteinShake: Building Datasets and Benchmarks for Deep Learning on Protein Structures Tim Kucera*, Carlos Oliver*, Dexiong Chen, Karsten Borgwardt NeurIPS Datasets and Benchmarks (2023) (preprint) |
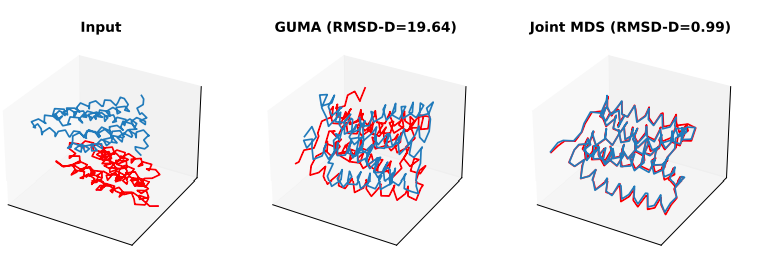 |
Unsupervised Manifold Alignment with Joint Multidimensional Scaling Dexiong Chen, Bowen Fan, Carlos Oliver, Karsten Borgwardt ICLR 2022 (preprint) |
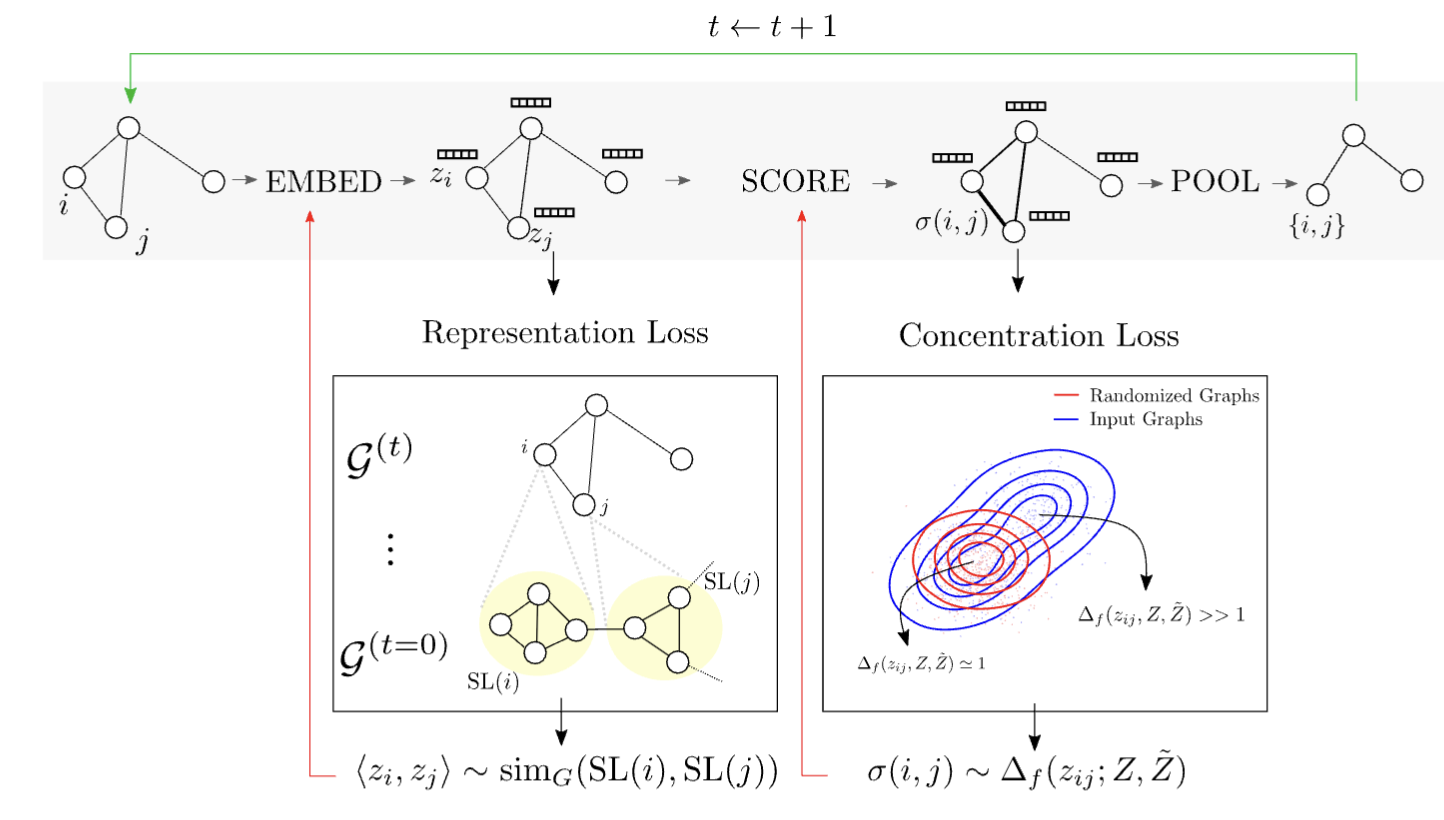 |
Approximate Network Motif Mining via Graph Learning Carlos Oliver, Dexiong Chen, Vincent Mallet, Pericles Philippopoulos, Karsten Borgwardt ArXiV (2022) (preprint) |
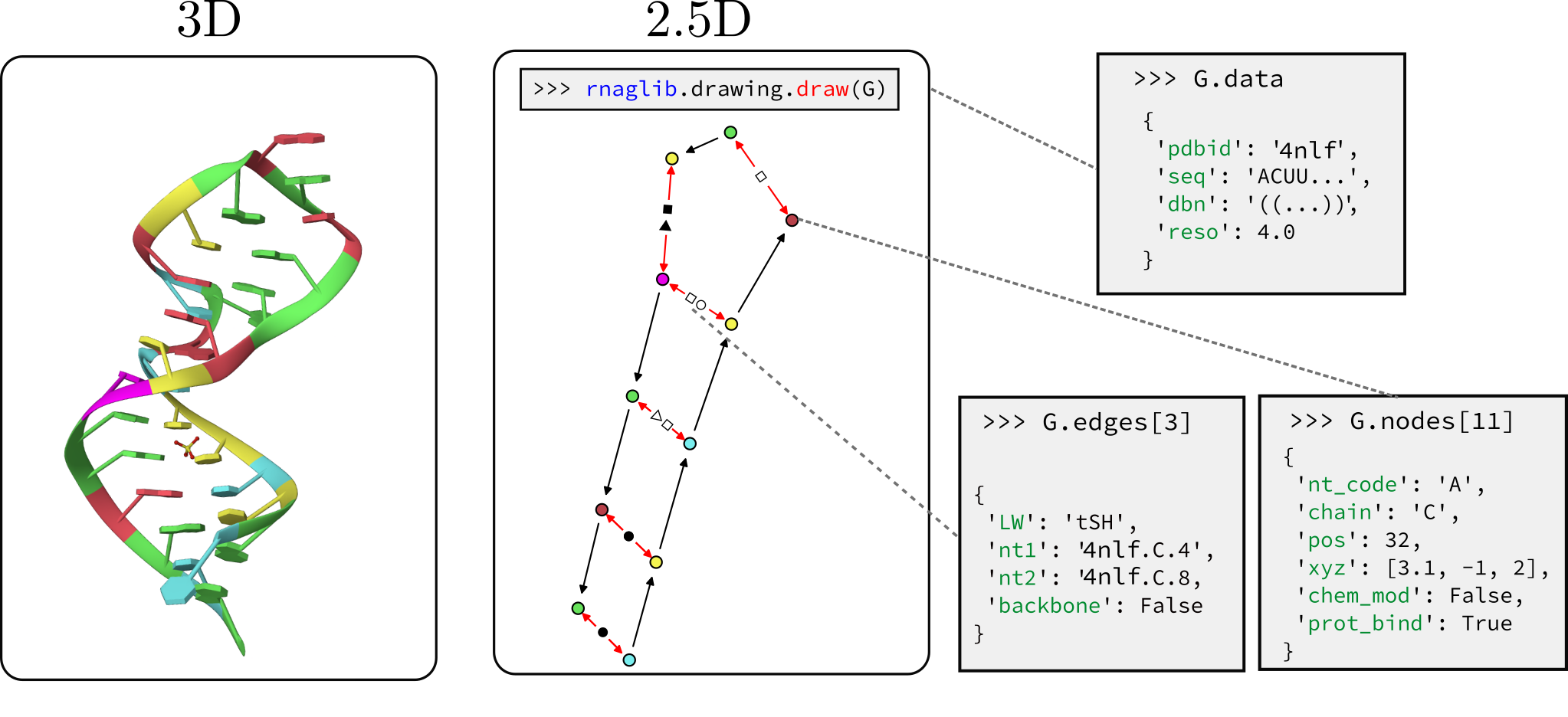 |
RNAGlib: A Python Package for RNA 2.5D Graphs Vincent Mallet, Carlos Oliver, Jonathan Broadbent, William L. Hamilton, Jerome Waldispuhl Bioinformatics (2021) (preprint | article) |
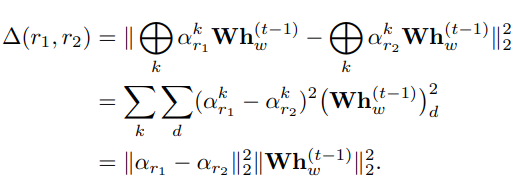 |
Edge Similarity-Aware Graph Neural Networks Vincent Mallet, Carlos Oliver, William L. Hamilton arXiv (2021) (preprint) |
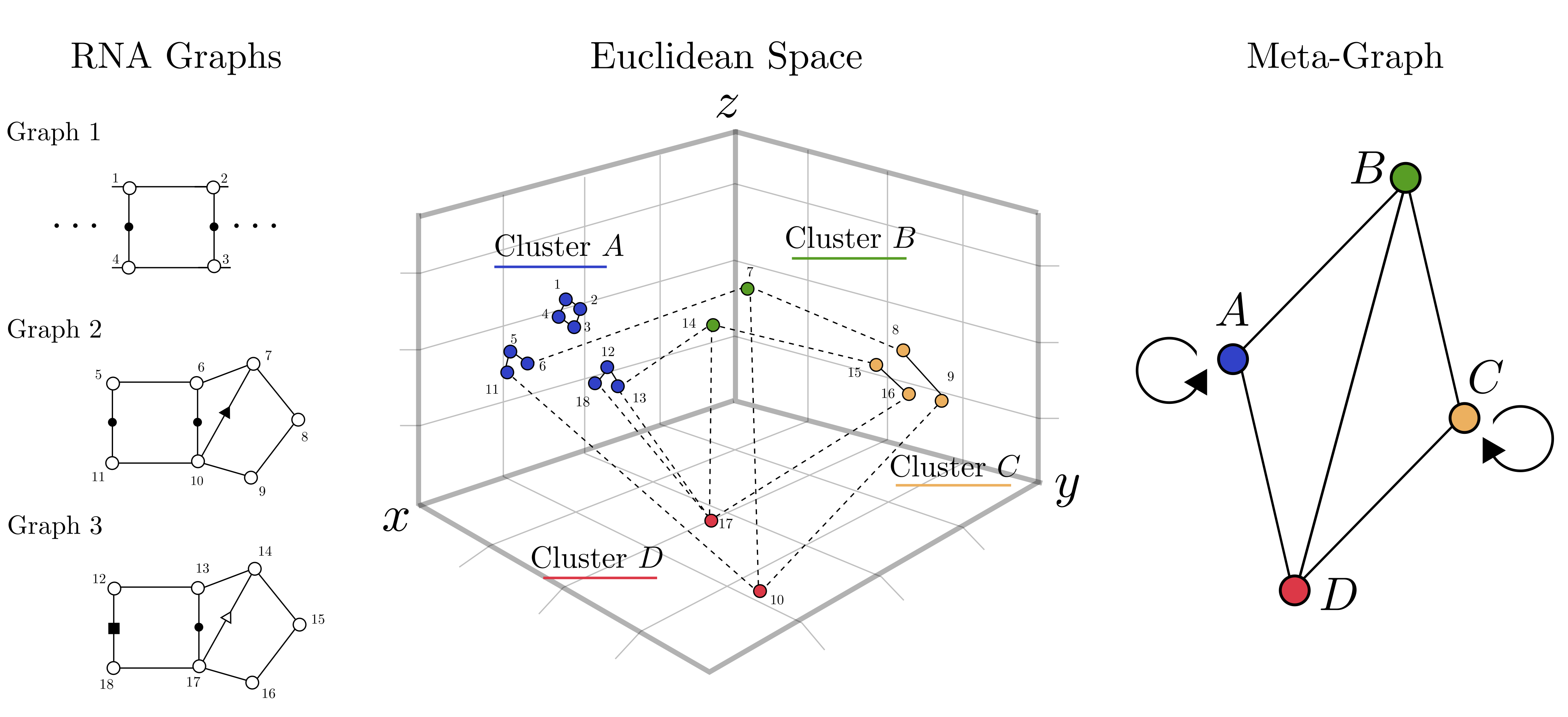 |
VeRNAl: A tool for fuzzy network motif mining in RNA Carlos Oliver, Vincent Mallet, Pericles Philippopoulos, William L. Hamilton, Jerome Waldispuhl RECOMB 2021 & Bioinformatics (preprint | article) |
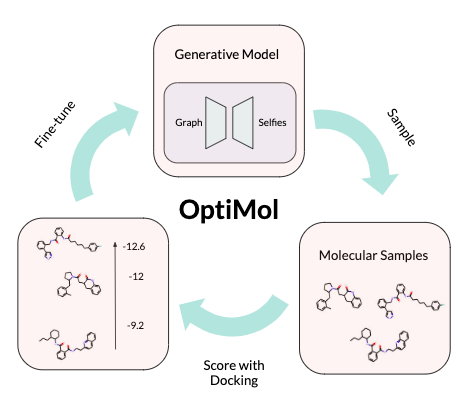 |
OptiMol: Optimization of binding affinities in chemical space for drug discovery Jacques Boitreaud, Vincent Mallet, Carlos Oliver, Jerome Waldispuhl ACM JCIM (2020) (article | preprint) |
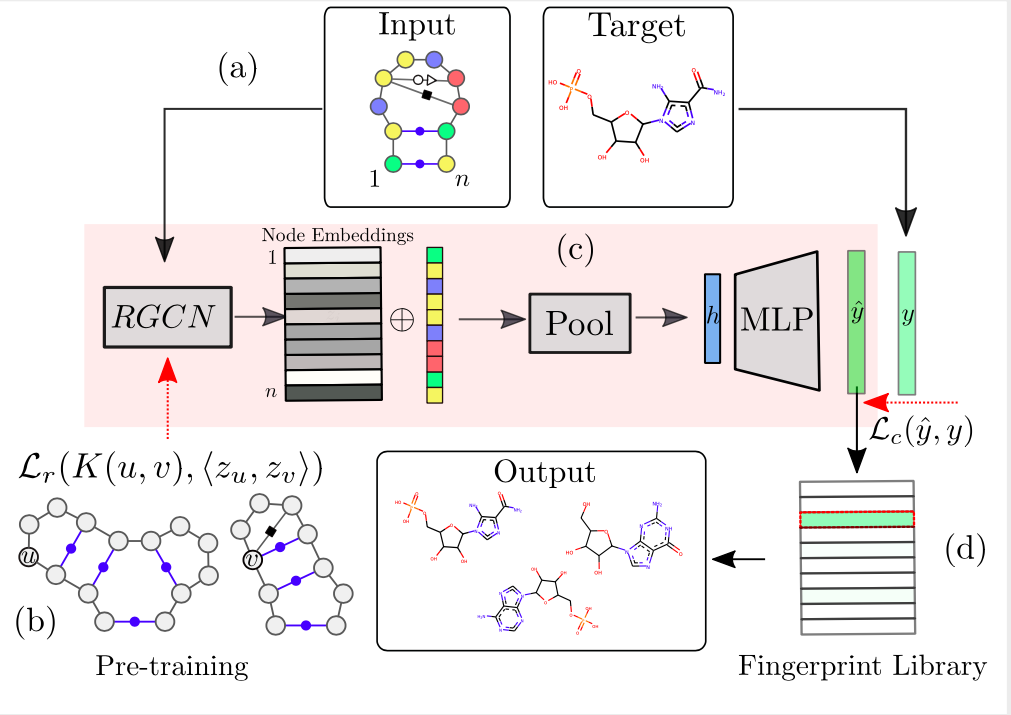 |
Augmented RNA base pairing networks imprint small molecule binding preferences Carlos Oliver, Vincent Mallet, Roman Sarrazin Gendron, Vladimir Reinharz, William L. Hamilton, Nicolas Moitessier, Jerome Waldispuhl Nucleic Acids Research (2020) (article | preprint) |
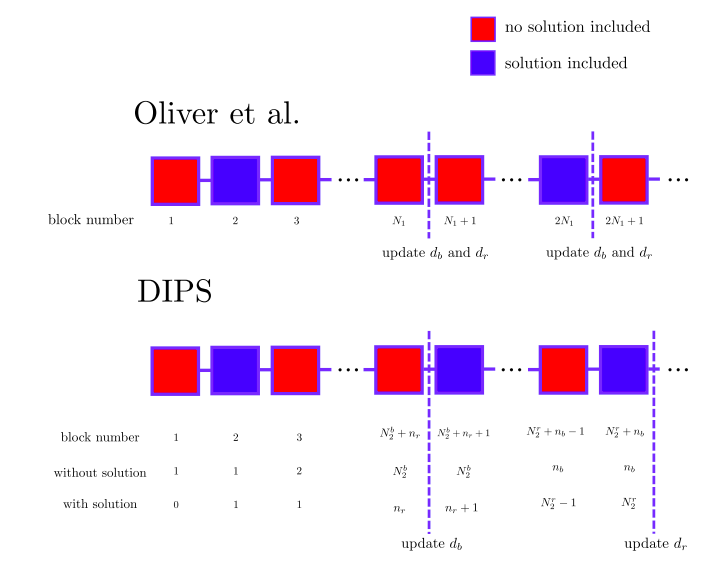 |
Difficulty Scaling in Proof of Work for Decentralized Problem Solving Pericles Philippopoulos, Alessandro Ricottone, Carlos Oliver Ledger Journal (2020) (article) |
 |
Stochastic Sampling of Structural Contexts Improves the Scalability and Accuracy of RNA 3D Module Identification Roman Sarrazin Gendron, Hua-Ting Yao, Vladimir Reinharz, Carlos Oliver, Yann Pony, Jerome Waldispuhl Accepted at RECOMB 2020 (preprint) |
| <img src="/assets/tarlig.png" class |