Research Group
This is regarding the direction of my project research group the Max Planck Institute for Biochemistry.
The spatial arrangement in 3D (structure) of biomolecules dictates what kinds of physical interactions are favorable and therefore which biological functions are accessible to DNA/RNA/proteins. Finding exactly which geometrical patterns within these molecules are the ones we should pay attention to is a needle in the haystack problem. In this group, we take the data-driven approach. That is, we build scalable structure-aware algorithms that process the growing datasets of biological structures to identify patterns which influence function. A clear understanding of the patterns that mediate structure to function relationships will help uncover new biological mechanisms and point towards solutions to pathologies.
Here are some active projects.
Deep learning on protein structures
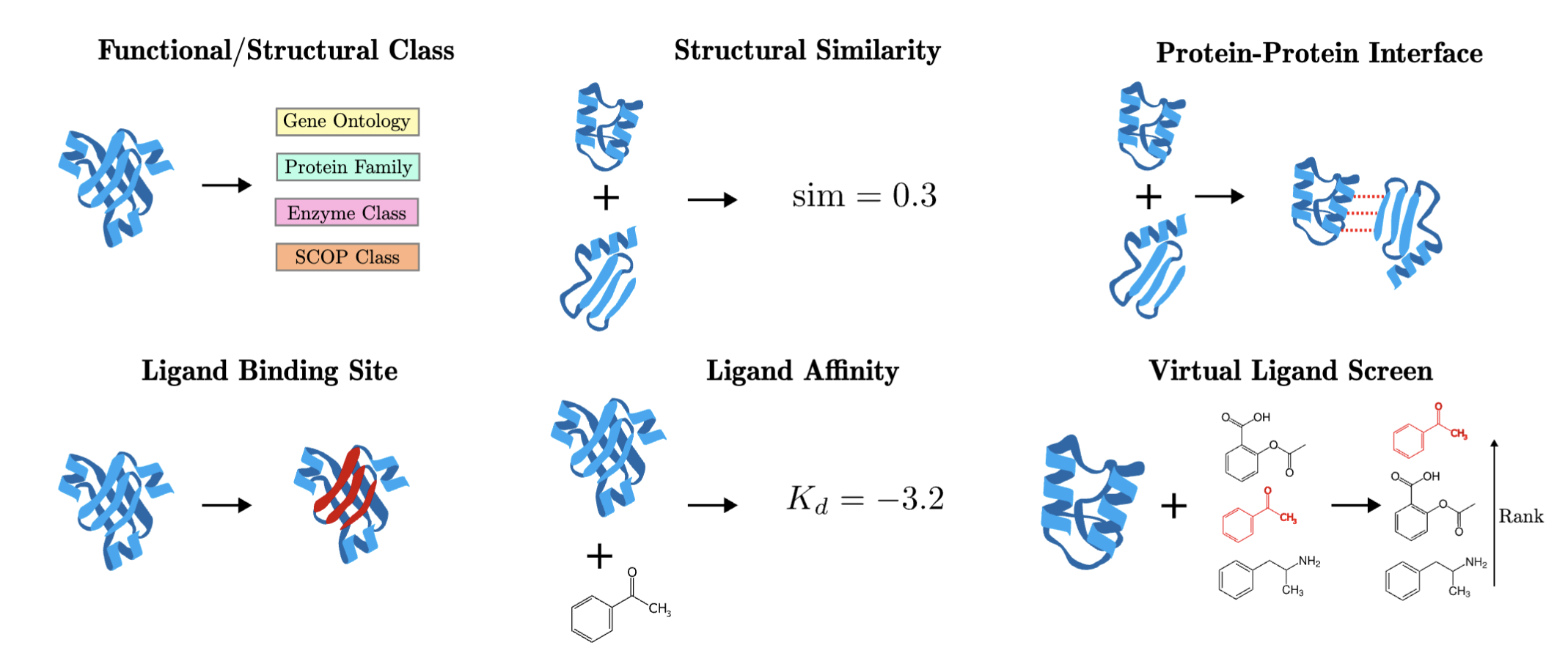
We recently developed ProteinShake which allows deep learning practitioners easy asccess to impactful biological tasks on proteins. (Accepted at NeurIPS 2023)
Representation learning on RNA 3D networks
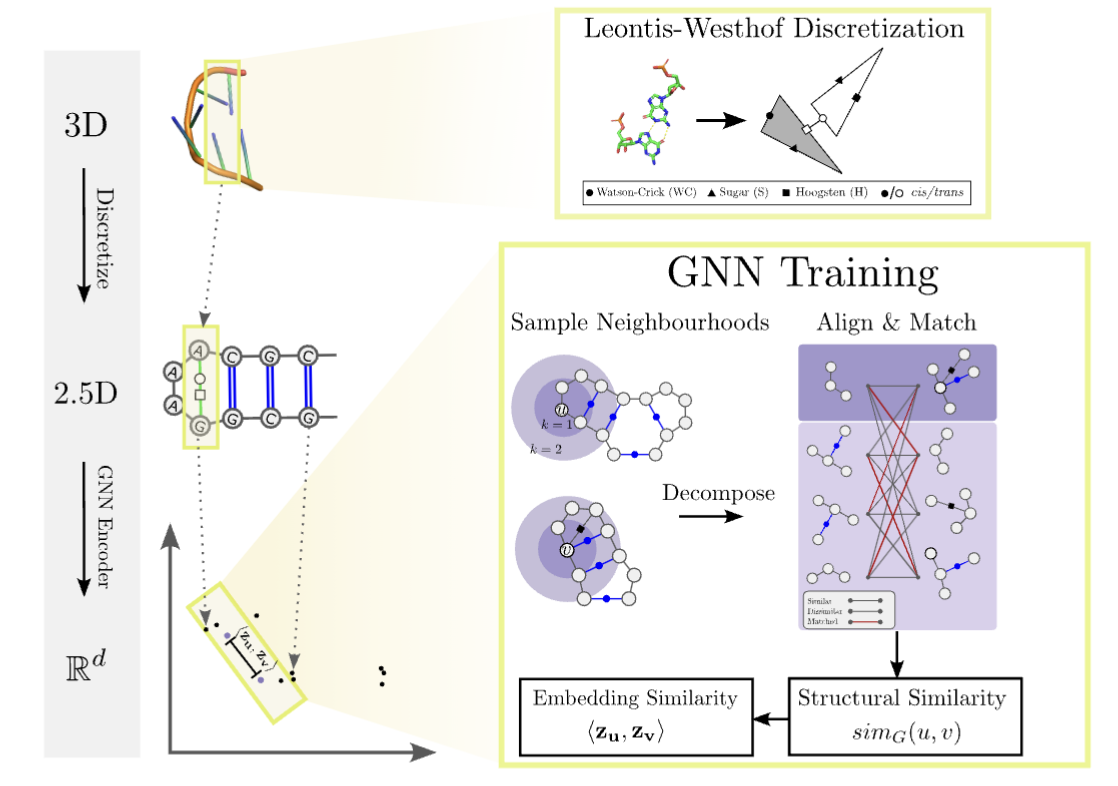
RNA structures adopt complex folds by forming base pairs, we have developed unsupervised learning methods which have proven useful in drug discovery and network motif mining applications. Through rnaglib we hope to explore more applications.
Graph pattern mining
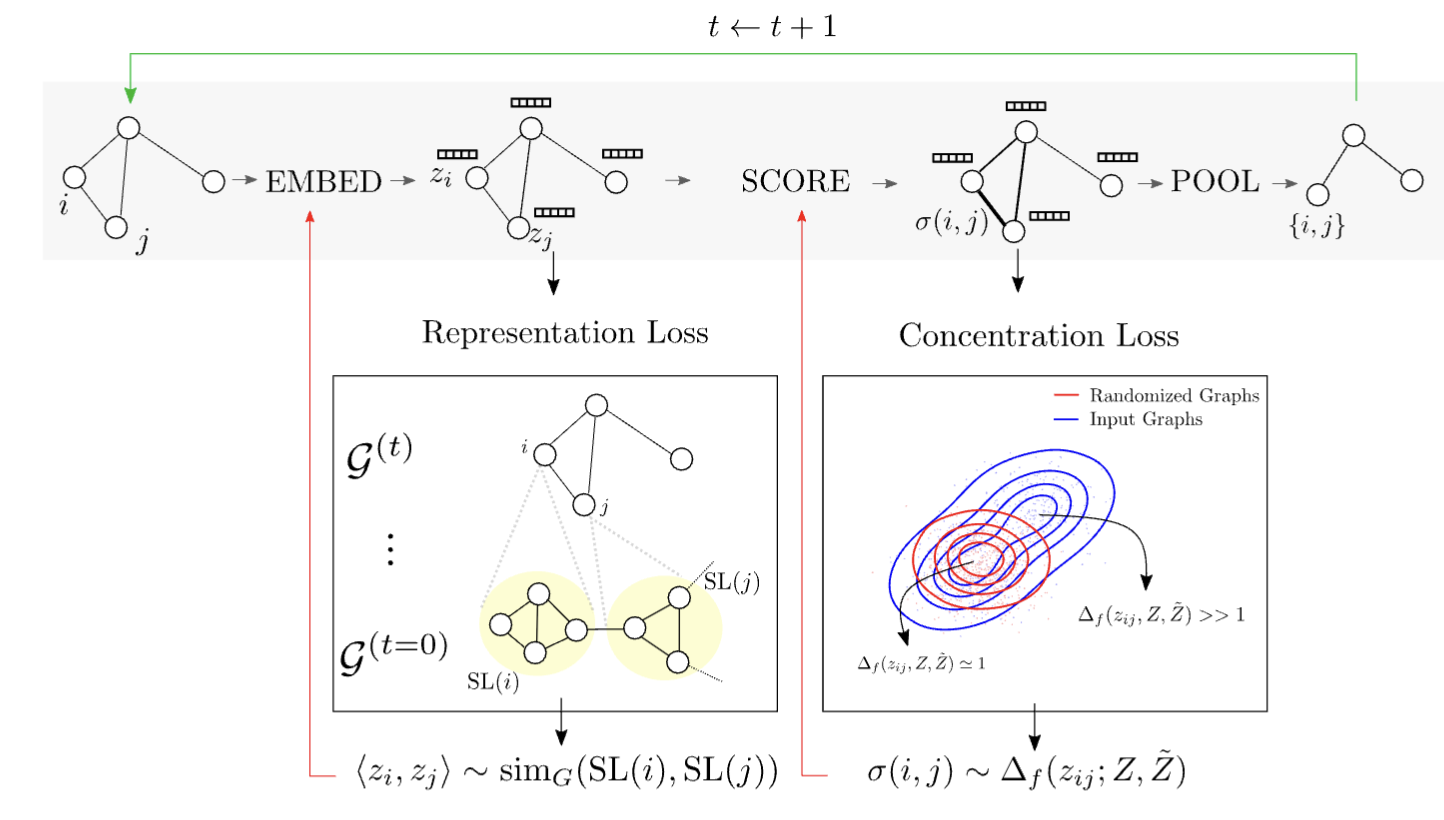
Structural patterns adopted by molecules are often dynamic and noisy, making traditional graph pattern mining algorithms less appropriate. MotiFiesta captures approximate network patterns in any graph dataset while at the same time serving as pre-training for classification models.
Open Positions
If you find this interesting, we are looking for a highly motivated PhD student to work at the intersection of structural biology, deep learning and pattern mining on graphs. (posting)(submit application)
Please contact me for more info: oliver@biochem.mpg.de